To make the most out of the features offered by the ASTR package,
input data should be imported into ASTR objects.
ASTR objects represent archaeometric and geochemical data
in R in a standardized and semantically rich format, to allow for
convenient and safe data analysis. If the input data structure adheres
to the conventions set out in the ASTR schema, then the package can
automatically build valid ASTR objects from it. When
working with the package, many of its functions require only an
ASTR object, making them convenient to use even for users
who consider themselves less experienced with R. This article highlights
some of these functionalities.
Before we start, we want to load the ASTR package:
Data import and inspection
To work with an ASTR object, we have to create one. The
function read_ASTR() imports spreadsheets in a variety of
formats, including Excel files. For this article, we use mock-up
data:
#> Group Cu_wt% Cu_err2SD% Sn_wt% As_wt% Sb_wt% Ag_ppm Ni_ppm d65Cu 206Pb/204Pb
#> 1 A 87.9 0.30 9.4 2.46 0.57 111 533 0.25 NA
#> 2 A 92.9 0.25 8.5 2.40 0.71 211 333 0.13 NA
#> 3 A 89.1 0.27 8.2 2.19 0.71 300 434 0.21 NA
#> 4 A 93.8 0.26 10.0 2.30 0.68 130 396 -0.5 NA
#> 5 A 94.4 0.21 8.3 1.52 0.58 240 585 -0.02 NA
#> 6 B 85.5 0.29 8.5 1.98 0.57 152 457 -0.28 18.4899
#> 7 B 90.3 0.22 7.7 2.26 0.62 132 584 #REF! 18.6064
#> 8 B 93.9 0.20 8.0 1.72 0.73 288 584 #REF! 18.4759
#> 9 B 90.5 0.23 6.4 1.82 0.63 324 578 #REF! 18.3556
#> 10 B 89.6 0.30 5.7 1.73 0.93 194 454 #REF! 18.4218
#> 206Pb/204Pb_err2SD 207Pb/204Pb 207Pb/204Pb_err2SD 208Pb/204Pb
#> 1 NA NA NA NA
#> 2 NA NA NA NA
#> 3 NA NA NA NA
#> 4 NA NA NA NA
#> 5 NA NA NA NA
#> 6 0.00097 15.6985 0.00080 38.391
#> 7 0.00087 15.6893 0.00070 38.047
#> 8 0.00102 15.6886 0.00080 38.233
#> 9 0.00074 15.6175 0.00070 38.256
#> 10 0.00087 15.6131 0.00066 38.300
#> 208Pb/204Pb_err2SD
#> 1 NA
#> 2 NA
#> 3 NA
#> 4 NA
#> 5 NA
#> 6 0.0013
#> 7 0.0015
#> 8 0.0020
#> 9 0.0015
#> 10 0.0019Note how the column headers are organised according to the ASTR
conventions. This allows the package to understand some of the
semantics of the dataset in the read_ASTR() process, and
correctly represent them in the resulting ASTR object. A
closer look at the mock-up data also reveals (common) problems with the
copper isotope data: it seems that some Excel formulas did not work as
intended and left #REF! entries. read_ASTR()
will read these as NA values. Finally, every
ASTR object requires one column that acts as a unique row
identifier. In this dataset there is no such column, though, only a
Group identifier. read_ASTR() can
automatically turn such a column to a unique identifier.
We can now perform the reading process. As we are working with
mock-up data in an R vignette, we do not read from the file system. We
only need to call an essential internal function of
read_ASTR(): as_ASTR(). It turns R data.frames
to ASTR objects. If we would start from an Excel file we
would call read_ASTR() instead.
To do this in practice, we not only have to submit our
data to as_ASTR(), but also set two other
arguments: 1. We have to define one of the columns as the ID column with
id_column, and 2. we explicitly have to mark any column
providing contextual information with context (i.e. no
analytical values, cf. ASTR
schema: Implementation).
data <- as_ASTR(data, id_column = "Group", context = c("Group"))
#> Warning in as_ASTR(data, id_column = "Group", context = c("Group")): Detected
#> multiple data rows with the same ID. They will be renamed consecutively using
#> the following convention: _1, _2, ... _n
#> Warning: Issue when transforming column "d65Cu" to numeric values: NAs
#> introduced by coercion
#> Warning in validate.ASTR(df6, quiet = FALSE): 34 missing values across 7
#> analytical columns
#> Warning in as_ASTR(data, id_column = "Group", context = c("Group")): See the
#> full list of validation output with: ASTR::validate(<your ASTR object>).While ASTR attempts to make data import as easy as possible, it also
aims to be transparent about it. Consequently, as_ASTR()
issues several warnings when confronted with our mock-up data, flagging
(potential) issues:
- The column for the ID did contain non-unique values but ASTR automatically created unique values from it.
- Some values in the column
d65Cuwere replaced withNA; when inspecting the data set, it is clear that these were the Excel#REF!error messages. In general, ASTR replaces by default common and user-defined indicators for missing values and also for indicators of “below detection limit” during import. - Across the dataset, 34 values are missing.
To learn more about the (potential) issues with our dataset, it
suggests to run validate():
validate(data)
#> # A tibble: 7 × 3
#> column count warning
#> <chr> <int> <chr>
#> 1 d65Cu 4 missing values
#> 2 206Pb/204Pb 5 missing values
#> 3 206Pb/204Pb_err2SD 5 missing values
#> 4 207Pb/204Pb 5 missing values
#> 5 207Pb/204Pb_err2SD 5 missing values
#> 6 208Pb/204Pb 5 missing values
#> 7 208Pb/204Pb_err2SD 5 missing valuesThis summary of where to find the missing values comes in handy for large datasets!
In our case, the dataset is fine. We might want to check on the missing values in the copper isotope data because apparently there was something wrong with the Excel file. In case of the lead isotope data, however, they are intentional (they were not measured for samples from group A).
Having a closer look on the imported data set shows, that ASTR did two more things:
data
#> ASTR table
#> Analytical columns: Cu, Cu_err2SD, Sn, As, Sb, Ag, Ni, d65Cu, 206Pb/204Pb, 206Pb/204Pb_err2SD, 207Pb/204Pb, 207Pb/204Pb_err2SD, 208Pb/204Pb, 208Pb/204Pb_err2SD
#> Contextual columns: Group
#> # A data frame: 10 × 16
#> ID Group Cu Cu_err2SD Sn As Sb Ag Ni d65Cu `206Pb/204Pb`
#> <chr> <chr> [wtP] [wtP] [wtP] [wtP] [wtP] [mg/… [mg/… <dbl> <dbl>
#> 1 A_1 A 87.9 0.264 9.4 2.46 0.57 111 533 0.25 NA
#> 2 A_2 A 92.9 0.232 8.5 2.4 0.71 211 333 0.13 NA
#> 3 A_3 A 89.1 0.241 8.2 2.19 0.71 300 434 0.21 NA
#> 4 A_4 A 93.8 0.244 10 2.3 0.68 130 396 -0.5 NA
#> 5 A_5 A 94.4 0.198 8.3 1.52 0.58 240 585 -0.02 NA
#> 6 B_1 B 85.5 0.248 8.5 1.98 0.57 152 457 -0.28 18.5
#> 7 B_2 B 90.3 0.199 7.7 2.26 0.62 132 584 NA 18.6
#> 8 B_3 B 93.9 0.188 8 1.72 0.73 288 584 NA 18.5
#> 9 B_4 B 90.5 0.208 6.4 1.82 0.63 324 578 NA 18.4
#> 10 B_5 B 89.6 0.269 5.7 1.73 0.93 194 454 NA 18.4
#> # ℹ 5 more variables: `206Pb/204Pb_err2SD` <dbl>, `207Pb/204Pb` <dbl>,
#> # `207Pb/204Pb_err2SD` <dbl>, `208Pb/204Pb` <dbl>, `208Pb/204Pb_err2SD` <dbl>The relative analytical precisions of the copper concentrations were
automatically converted to absolute analytical precisions, facilitating,
e.g., calculations and plotting. And our relative unit “ppm” was
converted into SI-units; the values now have a measurement unit R can
understand (note that the unit wt% is written as
wtP)1:
units(data$Ag[1])
#> $numerator
#> [1] "mg"
#>
#> $denominator
#> [1] "kg"
#>
#> attr(,"class")
#> [1] "symbolic_units"When looking into the data structure, we also see that it also labelled the different columns as e.g. concentration or isotope ratio:
str(data)
#> ASTR [10 × 16] (S3: ASTR/tbl_df/tbl/data.frame)
#> $ ID : chr [1:10] "A_1" "A_2" "A_3" "A_4" ...
#> ..- attr(*, "ASTR_class")= chr "ASTR_id"
#> $ Group : chr [1:10] "A" "A" "A" "A" ...
#> ..- attr(*, "ASTR_class")= chr "ASTR_context"
#> $ Cu : Units: [wtP] num [1:10] 87.9 92.9 89.1 93.8 94.4 85.5 90.3 93.9 90.5 89.6
#> ..- attr(*, "ASTR_class")= chr "ASTR_concentration"
#> $ Cu_err2SD : Units: [wtP] num [1:10] 0.264 0.232 0.241 0.244 0.198 ...
#> ..- attr(*, "ASTR_class")= chr "ASTR_error"
#> $ Sn : Units: [wtP] num [1:10] 9.4 8.5 8.2 10 8.3 8.5 7.7 8 6.4 5.7
#> ..- attr(*, "ASTR_class")= chr "ASTR_concentration"
#> $ As : Units: [wtP] num [1:10] 2.46 2.4 2.19 2.3 1.52 1.98 2.26 1.72 1.82 1.73
#> ..- attr(*, "ASTR_class")= chr "ASTR_concentration"
#> $ Sb : Units: [wtP] num [1:10] 0.57 0.71 0.71 0.68 0.58 0.57 0.62 0.73 0.63 0.93
#> ..- attr(*, "ASTR_class")= chr "ASTR_concentration"
#> $ Ag : Units: [mg/kg] num [1:10] 111 211 300 130 240 152 132 288 324 194
#> ..- attr(*, "ASTR_class")= chr "ASTR_concentration"
#> $ Ni : Units: [mg/kg] num [1:10] 533 333 434 396 585 457 584 584 578 454
#> ..- attr(*, "ASTR_class")= chr "ASTR_concentration"
#> $ d65Cu : num [1:10] 0.25 0.13 0.21 -0.5 -0.02 -0.28 NA NA NA NA
#> ..- attr(*, "ASTR_class")= chr [1:2] "ASTR_isotope" "ASTR_ratio"
#> $ 206Pb/204Pb : num [1:10] NA NA NA NA NA ...
#> ..- attr(*, "ASTR_class")= chr [1:2] "ASTR_isotope" "ASTR_ratio"
#> $ 206Pb/204Pb_err2SD: num [1:10] NA NA NA NA NA 0.00097 0.00087 0.00102 0.00074 0.00087
#> ..- attr(*, "ASTR_class")= chr "ASTR_error"
#> $ 207Pb/204Pb : num [1:10] NA NA NA NA NA ...
#> ..- attr(*, "ASTR_class")= chr [1:2] "ASTR_isotope" "ASTR_ratio"
#> $ 207Pb/204Pb_err2SD: num [1:10] NA NA NA NA NA 0.0008 0.0007 0.0008 0.0007 0.00066
#> ..- attr(*, "ASTR_class")= chr "ASTR_error"
#> $ 208Pb/204Pb : num [1:10] NA NA NA NA NA ...
#> ..- attr(*, "ASTR_class")= chr [1:2] "ASTR_isotope" "ASTR_ratio"
#> $ 208Pb/204Pb_err2SD: num [1:10] NA NA NA NA NA 0.0013 0.0015 0.002 0.0015 0.0019
#> ..- attr(*, "ASTR_class")= chr "ASTR_error"Subsetting data
With this much information about our data, it is easy to subset data.
Let’s say that, for example, we want to check if our data can be
differentiated into groups based on the chemical composition by creating
biplots of all possible combinations. The
get_ ... _columns()-family comes in very handy here:
pairs(get_concentration_columns(data)[-1])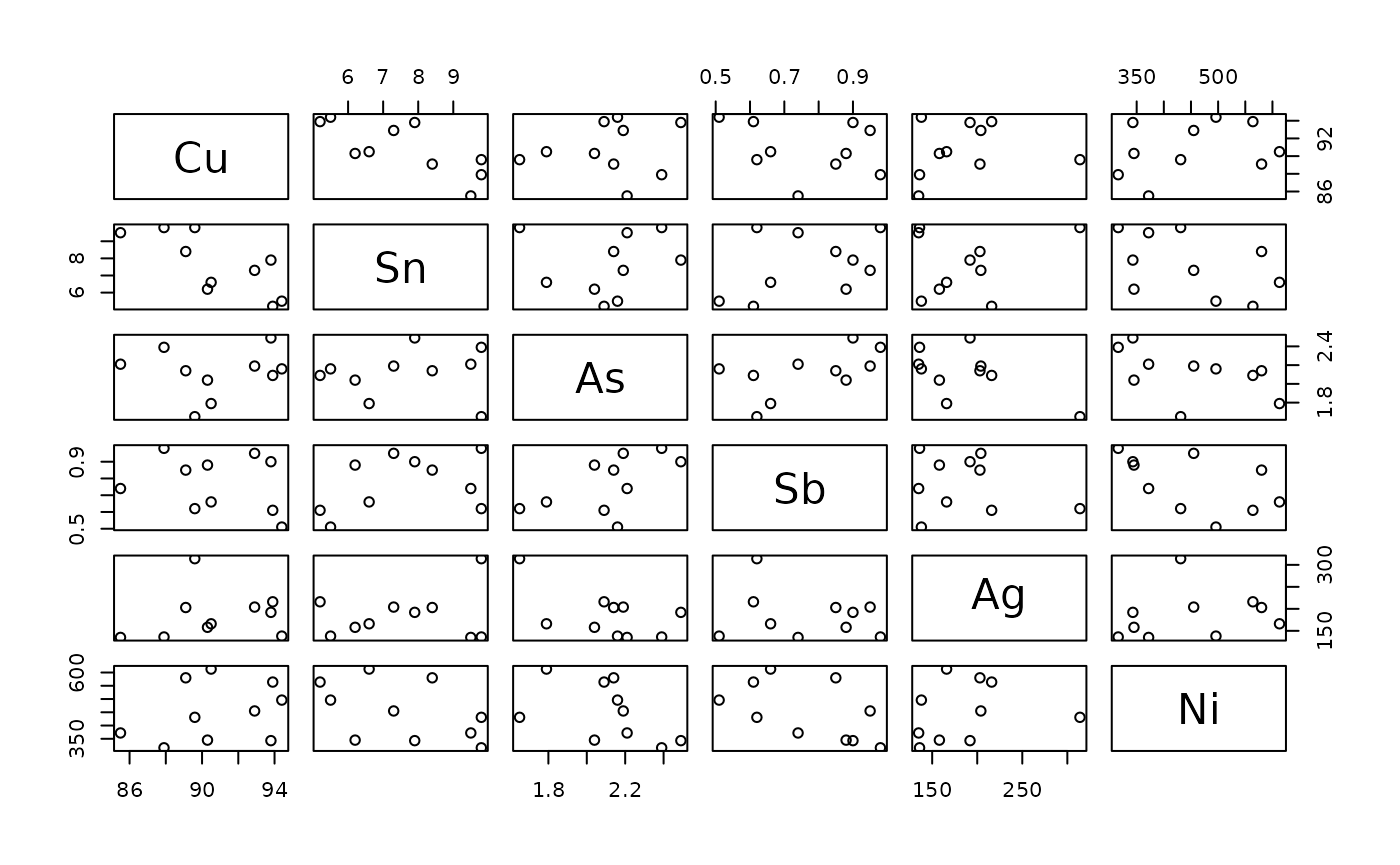
Note that we have to exclude the first column here, because the first
column of ASTR objects and their subsets is always the ID
column to ensure that data can be clearly assigned to a single sample.
Actually, we might only be interested in the main elements of the metal.
In this case, we can take advantage that these are the only values in
wtP:
pairs(get_unit_columns(data, units = "wtP")[-1])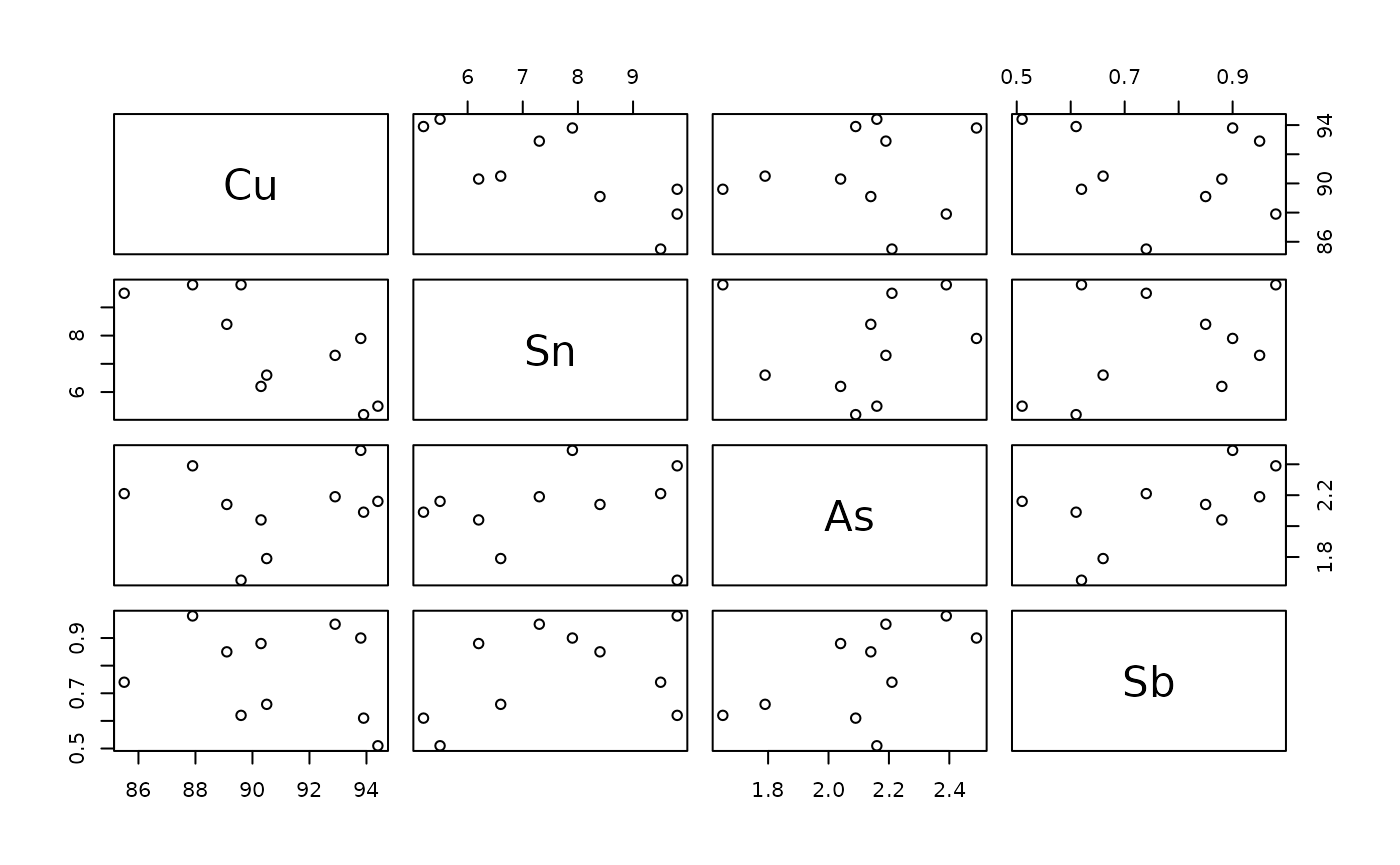
Unit conversions
While continuing investigating the metal, we may want to estimate its
approximate melting temperature and remember that we can get this
information from a phase diagram. While our values for copper and tin
are in wt%, the phase diagram in front of us is in
at%. Converting from wt% to at%
is probably faster than searching for a version of the phase diagram in
wt%2:
library(magrittr) # import of pipe operator `%>%`
unify_concentration_unit(data, unit = "wtP") %>%
wt_to_at()
#> ASTR table
#> Analytical columns: Cu, Cu_err2SD, Sn, As, Sb, Ag, Ni, d65Cu, 206Pb/204Pb, 206Pb/204Pb_err2SD, 207Pb/204Pb, 207Pb/204Pb_err2SD, 208Pb/204Pb, 208Pb/204Pb_err2SD
#> Contextual columns: Group
#> # A data frame: 10 × 16
#> ID Group Cu Cu_err2SD Sn As Sb Ag Ni d65Cu
#> <chr> <chr> [atP] [wtP] [atP] [atP] [atP] [atP] [atP] <dbl>
#> 1 A_1 A 92.2 0.264 5.28 2.19 0.312 0.00686 0.0605 0.25
#> 2 A_2 A 93.0 0.232 4.55 2.04 0.371 0.0124 0.0361 0.13
#> 3 A_3 A 93.0 0.241 4.58 1.94 0.387 0.0185 0.0491 0.21
#> 4 A_4 A 92.4 0.244 5.27 1.92 0.350 0.00754 0.0422 -0.5
#> 5 A_5 A 93.9 0.198 4.42 1.28 0.301 0.0141 0.0630 -0.02
#> 6 B_1 B 92.8 0.248 4.94 1.82 0.323 0.00972 0.0537 -0.28
#> 7 B_2 B 93.3 0.199 4.26 1.98 0.335 0.00804 0.0654 NA
#> 8 B_3 B 93.8 0.188 4.28 1.46 0.381 0.0169 0.0632 NA
#> 9 B_4 B 94.4 0.208 3.57 1.61 0.343 0.0199 0.0653 NA
#> 10 B_5 B 94.7 0.269 3.22 1.55 0.513 0.0121 0.0519 NA
#> # ℹ 6 more variables: `206Pb/204Pb` <dbl>, `206Pb/204Pb_err2SD` <dbl>,
#> # `207Pb/204Pb` <dbl>, `207Pb/204Pb_err2SD` <dbl>, `208Pb/204Pb` <dbl>,
#> # `208Pb/204Pb_err2SD` <dbl>We use a two-step approach here to be more accurate: The actual
conversion from wt% to at% is done by
wt_to_at(). However, because Ag and Ni are not in
wt%, they need to be converted first. That’s why we run
unify_concentration_unit() first: it detects values with
units that can be converted into the target unit and converts them all
into the same unit. While it is nice to be scientifically accurate, we
can get only a rough estimate anyway because of the significant amounts
of arsenic and antimony in the metal. Therefore, converting only copper
and tin would be sufficient:
wt_to_at(data, elements = c("Cu", "Sn"))
#> ASTR table
#> Analytical columns: Cu, Cu_err2SD, Sn, As, Sb, Ag, Ni, d65Cu, 206Pb/204Pb, 206Pb/204Pb_err2SD, 207Pb/204Pb, 207Pb/204Pb_err2SD, 208Pb/204Pb, 208Pb/204Pb_err2SD
#> Contextual columns: Group
#> # A data frame: 10 × 16
#> ID Group Cu Cu_err2SD Sn As Sb Ag Ni d65Cu `206Pb/204Pb`
#> <chr> <chr> [atP] [wtP] [atP] [wtP] [wtP] [mg/… [mg/… <dbl> <dbl>
#> 1 A_1 A 94.6 0.264 5.41 2.46 0.57 111 533 0.25 NA
#> 2 A_2 A 95.3 0.232 4.67 2.4 0.71 211 333 0.13 NA
#> 3 A_3 A 95.3 0.241 4.70 2.19 0.71 300 434 0.21 NA
#> 4 A_4 A 94.6 0.244 5.40 2.3 0.68 130 396 -0.5 NA
#> 5 A_5 A 95.5 0.198 4.50 1.52 0.58 240 585 -0.02 NA
#> 6 B_1 B 94.9 0.248 5.05 1.98 0.57 152 457 -0.28 18.5
#> 7 B_2 B 95.6 0.199 4.37 2.26 0.62 132 584 NA 18.6
#> 8 B_3 B 95.6 0.188 4.36 1.72 0.73 288 584 NA 18.5
#> 9 B_4 B 96.4 0.208 3.65 1.82 0.63 324 578 NA 18.4
#> 10 B_5 B 96.7 0.269 3.29 1.73 0.93 194 454 NA 18.4
#> # ℹ 5 more variables: `206Pb/204Pb_err2SD` <dbl>, `207Pb/204Pb` <dbl>,
#> # `207Pb/204Pb_err2SD` <dbl>, `208Pb/204Pb` <dbl>, `208Pb/204Pb_err2SD` <dbl>Working with data
Another task in our investigation of the copper items is to identify the type of copper alloy our metal is made of. A first rough approach could be to use the classification suggested by Bray et al. (2015). Not surprising, we see that they all belong to the group “As+Sb”:
copper_group <- copper_group_bray(data)
copper_group
#> ASTR table
#> Analytical columns: As, Sb, Ag, Ni
#> Contextual columns: Group, copper_group_bray
#> # A data frame: 10 × 7
#> ID Group As Sb Ag Ni copper_group_bray
#> <chr> <chr> [wtP] [wtP] [wtP] [wtP] <chr>
#> 1 A_1 A 2.46 0.57 0.0111 0.0533 As+Sb
#> 2 A_2 A 2.4 0.71 0.0211 0.0333 As+Sb
#> 3 A_3 A 2.19 0.71 0.03 0.0434 As+Sb
#> 4 A_4 A 2.3 0.68 0.013 0.0396 As+Sb
#> 5 A_5 A 1.52 0.58 0.024 0.0585 As+Sb
#> 6 B_1 B 1.98 0.57 0.0152 0.0457 As+Sb
#> 7 B_2 B 2.26 0.62 0.0132 0.0584 As+Sb
#> 8 B_3 B 1.72 0.73 0.0288 0.0584 As+Sb
#> 9 B_4 B 1.82 0.63 0.0324 0.0578 As+Sb
#> 10 B_5 B 1.73 0.93 0.0194 0.0454 As+SbAfter we look into the chemical compositions, it is time to turn to the lead isotope data. Before plotting them, we want to calculate their age model parameters because they give us some idea about the geological setting. The model by Stacey & Kramers (1975) is still the most commonly used one. Because we have lead isotope data only for group B, we exclude data from group A:
data_pb_iso <- data[data$Group == "B", ]
age_model_params <- pb_iso_age_model(data_pb_iso, model = "SK75")
age_model_params
#> ASTR table
#> Analytical columns: 206Pb/204Pb, 207Pb/204Pb, 208Pb/204Pb
#> Contextual columns: Group, model_age_SK75, mu_SK75, kappa_SK75
#> # A data frame: 5 × 8
#> ID Group `206Pb/204Pb` `207Pb/204Pb` `208Pb/204Pb` model_age_SK75 mu_SK75
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 B_1 B 18.5 15.7 38.4 304. 10.1
#> 2 B_2 B 18.6 15.7 38.0 200. 10.0
#> 3 B_3 B 18.5 15.7 38.2 294. 10.1
#> 4 B_4 B 18.4 15.6 38.3 239. 9.77
#> 5 B_5 B 18.4 15.6 38.3 180. 9.73
#> # ℹ 1 more variable: kappa_SK75 <dbl>Data export
After finishing our analysis of the data, we want to save the copper
type and the parameters of the lead isotope model age to our hard drive.
For this task, ASTR relies on the functions already provided by R and
other R packages (e.g. readr::write_csv()). Before export,
we have to make sure that the information about the units of the values
is preserved by including it back in the column name:
data_unitless <- remove_units(data, recover_unit_names = TRUE)
data_unitless
#> ASTR table
#> Analytical columns: Cu_wt%, Cu_err2SD, Sn_wt%, As_wt%, Sb_wt%, Ag_mg/kg, Ni_mg/kg, d65Cu, 206Pb/204Pb, 206Pb/204Pb_err2SD, 207Pb/204Pb, 207Pb/204Pb_err2SD, 208Pb/204Pb, 208Pb/204Pb_err2SD
#> Contextual columns: Group
#> # A data frame: 10 × 16
#> ID Group `Cu_wt%` Cu_err2SD `Sn_wt%` `As_wt%` `Sb_wt%` `Ag_mg/kg`
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 A_1 A 87.9 0.264 9.4 2.46 0.57 111
#> 2 A_2 A 92.9 0.232 8.5 2.4 0.71 211
#> 3 A_3 A 89.1 0.241 8.2 2.19 0.71 300
#> 4 A_4 A 93.8 0.244 10 2.3 0.68 130
#> 5 A_5 A 94.4 0.198 8.3 1.52 0.58 240
#> 6 B_1 B 85.5 0.248 8.5 1.98 0.57 152
#> 7 B_2 B 90.3 0.199 7.7 2.26 0.62 132
#> 8 B_3 B 93.9 0.188 8 1.72 0.73 288
#> 9 B_4 B 90.5 0.208 6.4 1.82 0.63 324
#> 10 B_5 B 89.6 0.269 5.7 1.73 0.93 194
#> # ℹ 8 more variables: `Ni_mg/kg` <dbl>, d65Cu <dbl>, `206Pb/204Pb` <dbl>,
#> # `206Pb/204Pb_err2SD` <dbl>, `207Pb/204Pb` <dbl>,
#> # `207Pb/204Pb_err2SD` <dbl>, `208Pb/204Pb` <dbl>, `208Pb/204Pb_err2SD` <dbl>